Database statistics#
mOTUs taxonomy overview#
There are 124’295 mOTUs in release 4.1. Each mOTU is mapped on to the GTDB R226 taxonomy and summarised by taxonomic rank.
Bacteria |
Archaea |
Total |
|
|---|---|---|---|
Phylum |
145 |
19 |
164 |
Class |
410 |
52 |
462 |
Order |
1,431 |
115 |
1,546 |
Family |
3,653 |
370 |
4,023 |
Genus |
15,904 |
1,090 |
16,994 |
mOTU |
120,445 |
3,850 |
124,295 |
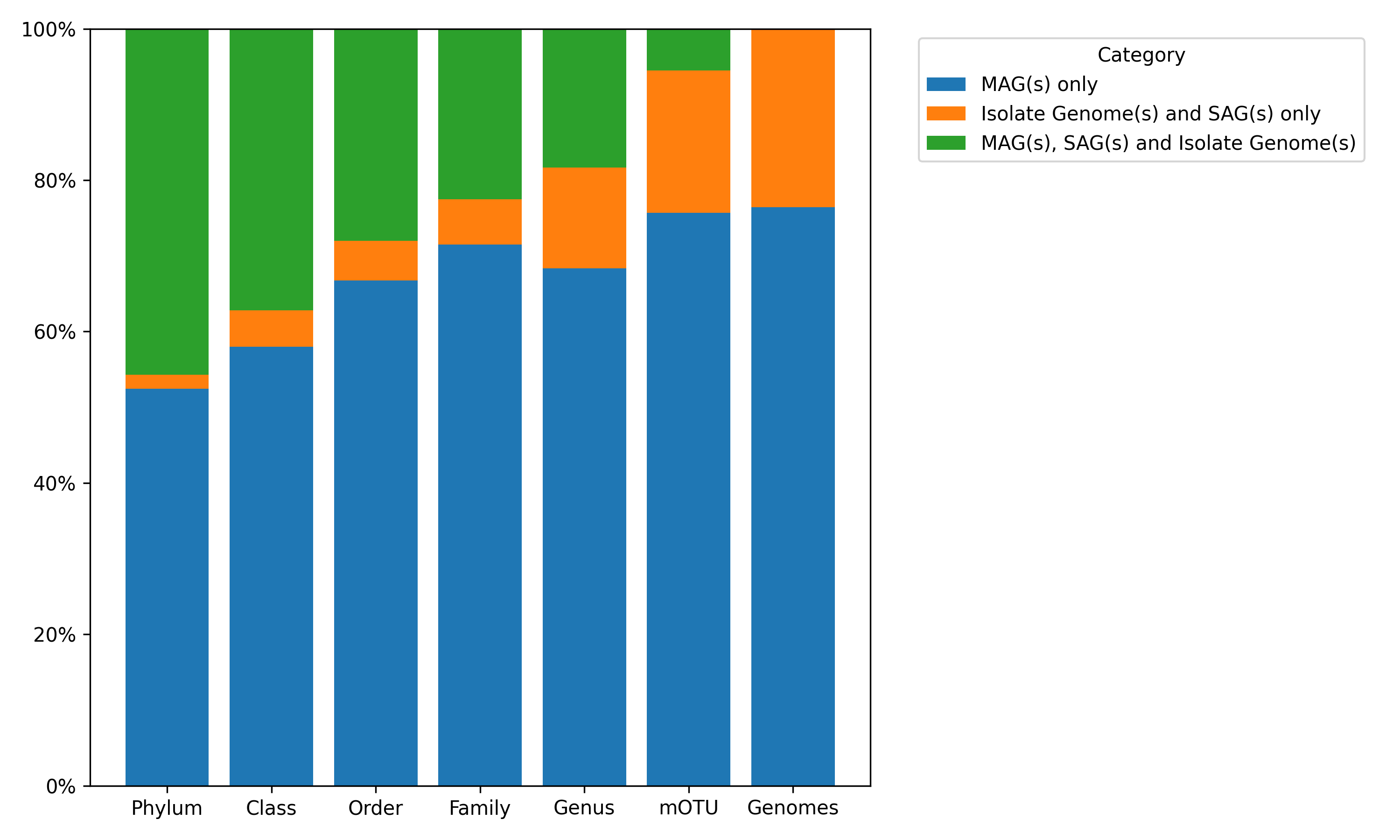
Genome Categories#
The mOTUs database contains ~3 million MAGs and ~1 million isolate/single-cell genomes, all mapped to the GTDB R226 taxonomy. The plot displays the percentage of taxonomic ranks represented by MAGs only, isolates only, or a combination of both (Mixed). The final “Genomes” bar reflects the total ratio of MAGs within the entire database.
Genome Quality#
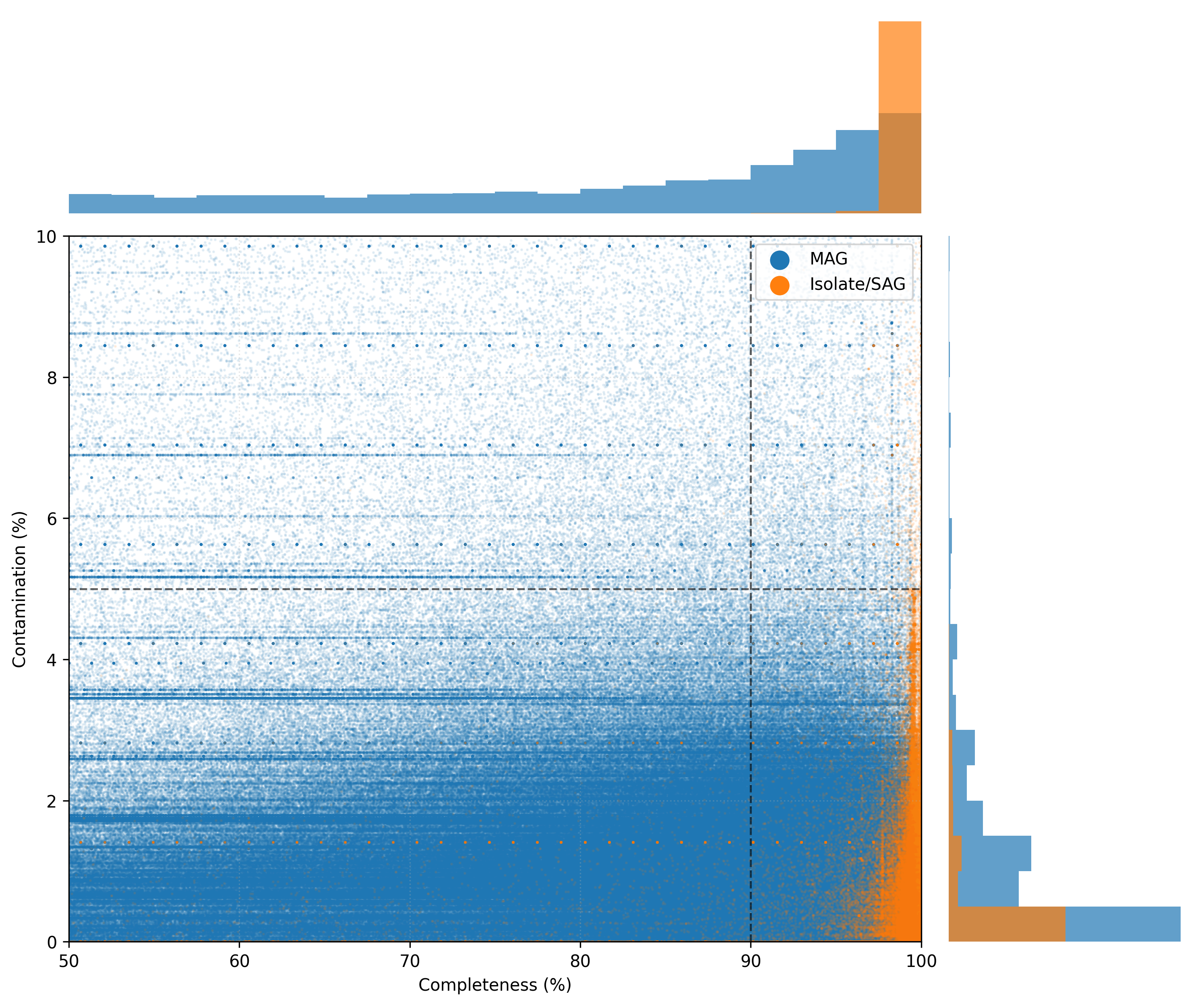
All genomes (~ 4 million) in the mOTUs 4.1 genomes database were quality controlled and bins with a completeness below 50% and completeness below 10% were removed (exceptions were made for Eremiobacterota, see here). The figure below shows the quality distribution for all genomes in the mOTUs 4.1 database.
Representative Genome Quality#
Representative genomes for each mOTU are selected based on a hierarchical scoring system designed to prioritize biological reliability and assembly quality.
Priority |
Criterion |
|---|---|
|
Isolate genomes and SAGs are preferred over MAGs. |
|
Highest Q-Score (Completeness - 5 * Contamination) |
|
Highest N50 (assembly statistic: half of the genome is contained in contigs of this length or longer). |
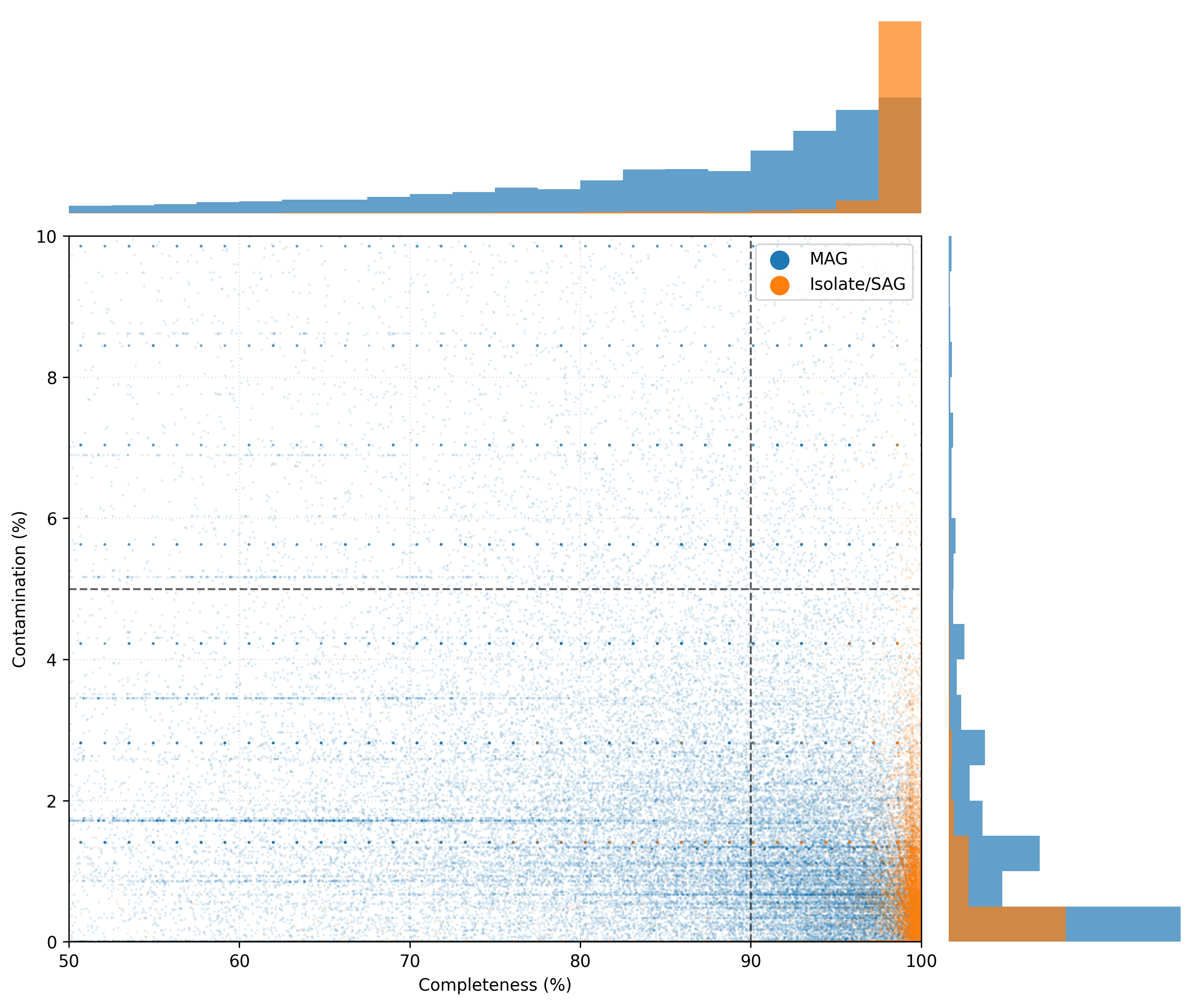
The figure below shows the distribution of genome quality of representative genomes:
Environmental annotation#
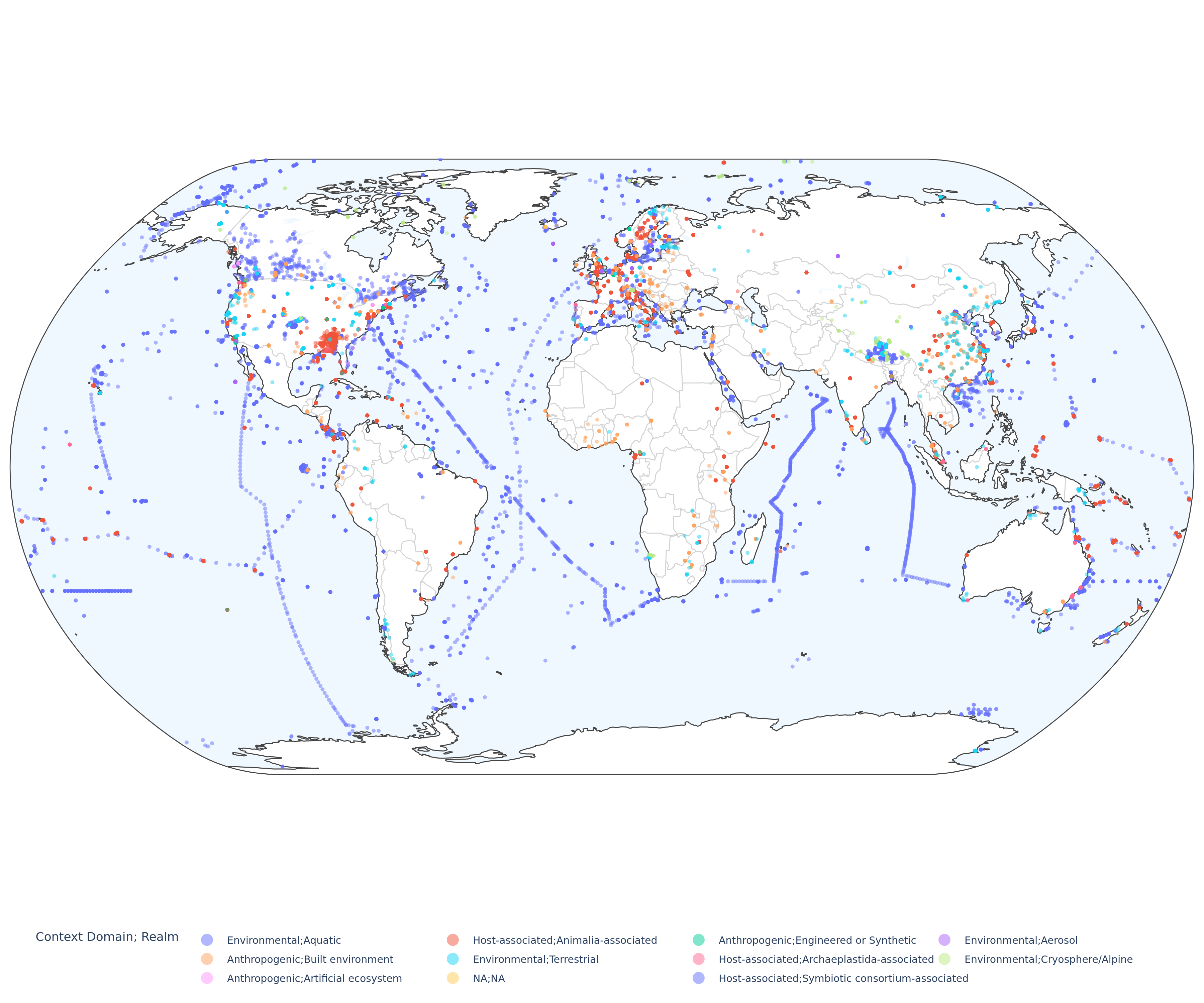
MAGs of the mOTUs 4.1 database were reconstructed from >120k short read metagenomic samples. Where possible samples were annotated using a hierarchical six level environmental ontology. The figure below shows the distribution of environments across samples.
Taxa with the largest number of mOTUs#
GTDB taxonomy of mOTUs sorted by number and taxonomic rank.
Phylum |
Class |
Order |
Family |
Genus |
|---|---|---|---|---|
Pseudomonadota (30’707) |
Clostridia (19’431) |
Bacteroidales (8’738) |
Lachnospiraceae (4’213) |
Streptococcus (2’185) |
Bacillota (29’895) |
Bacteroidia (17’706) |
Oscillospirales (8’353) |
Burkholderiaceae (3’782) |
Collinsella (1’807) |
Bacteroidota (17’706) |
Gammaproteobacteria (17’207) |
Burkholderiales (5’789) |
Flavobacteriaceae (3’085) |
Nanosyncoccus (1’541) |
Actinomycetota (13’272) |
Alphaproteobacteria (13’459) |
Flavobacteriales (4’962) |
Bacteroidaceae (2’887) |
Prevotella (1’455) |
Patescibacteriota (5’738) |
Bacilli (9’086) |
Lachnospirales (4’461) |
Oscillospiraceae (2’514) |
Cryptobacteroides (1’164) |
Verrucomicrobiota (3’097) |
Actinomycetes (8’175) |
Pseudomonadales (3’770) |
Streptococcaceae (2’224) |
Faecousia (1’028) |
Chloroflexota (2’882) |
Saccharimonadia (3’365) |
Saccharimonadales (3’327) |
Rhodobacteraceae (2’219) |
Flavobacterium (894) |
Cyanobacteriota (2’277) |
Coriobacteriia (2’926) |
Actinomycetales (3’243) |
CAG-272 (2’115) |
Pelagibacter (886) |
Planctomycetota (1’869) |
Verrucomicrobiia (2’154) |
Lactobacillales (3’175) |
Acutalibacteraceae (2’034) |
Streptomyces (734) |
Acidobacteriota (1’766) |
Acidimicrobiia (1’695) |
Christensenellales (2’943) |
UBA660 (1’943) |
Colicola (715) |
Global distribution of environmental samples#
 mOTUs is part of SIB's portfolio of open tools and databases.
mOTUs is part of SIB's portfolio of open tools and databases.
 mOTUs is part of the ELIXIR-CH Service Delivery Plan.
mOTUs is part of the ELIXIR-CH Service Delivery Plan.