mOTUs Tool#
In brief#
Microbial communities drive the cycling of energy, biomass, and nutrients on our planet, and are essential for the health of humans, animals, and plants. Taxonomic profiling of these communities - quantifying which taxa are present at what abundance, is a key step in identifying the drivers of microbial community structure, unraveling the complexity of microbial interactions, or discovering disease biomarkers.
The mOTUs command-line tool enables species-level taxonomic profiling of microbial communities from shotgun metagenomic sequencing data.
By using a set of universal single-copy marker genes (MGs), mOTUs is supported by, yet independent from, the availability of sequenced genomes
from isolated strains. As such, it enables reference genome-independent profiling of both known and unknown species.
The tool uses one or several FastQ/A-formatted metagenomic sequence file(s) as input and outputs a taxonomic
profile that reports the relative abundance of detected species either as read counts or proportions.
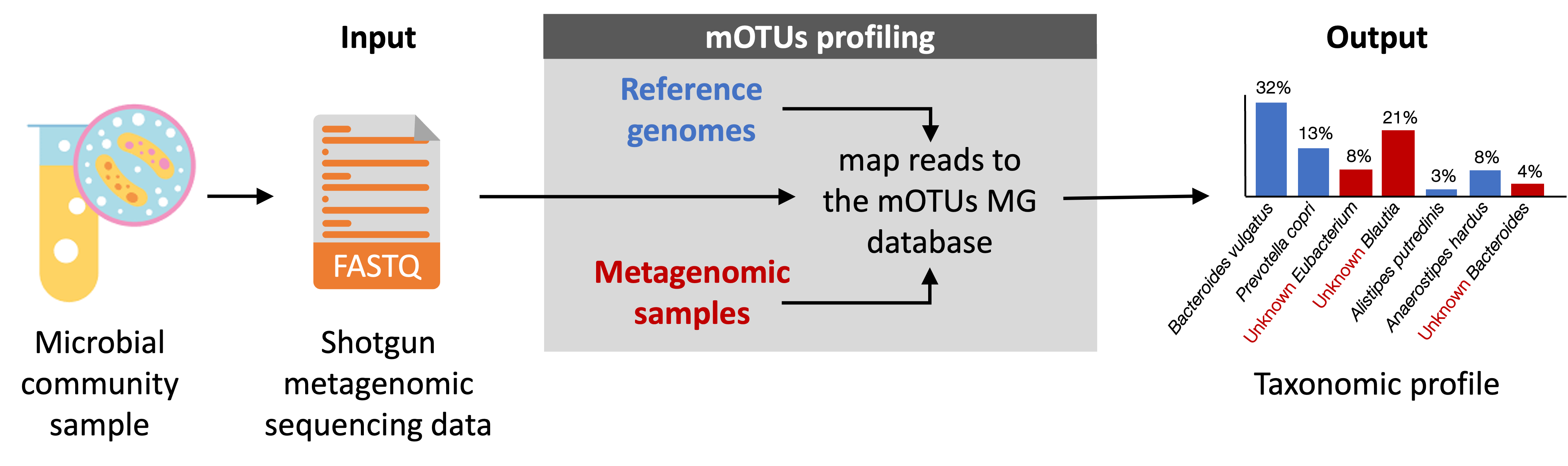
Why should you use mOTUs?#
🧬 Taxonomic Coverage: Profile >124k species
The mOTUs database is constructed from 919K isolate and single cell-amplified (SAGs) genomes available from reference databases and 2.97M metagenome-assembled genomes (MAGs) generated from 120,769 metagenomic samples spanning diverse microbiomes beyond the human gut or ocean microbiomes, for example, animal-associated microbiomes, freshwater, or wetlands, covering environments we found to be greatly underrepresented by reference genomes.
In the current version, 124,295 species-level taxonomic units (mOTUs) were constructed using sequences of 10 single-copy marker genes recovered from these genomes. 30,256 mOTUs are represented by an isolate genome, whereas 94,039 mOTUs are represented by MAGs only.
🎯 Precision Quantification: Identify and quantify species with high precision
Due to the compositional nature of metagenomic data, methods that rely on quantifying only species for which reference genomes are available produce biased results in which the relative abundance of such species in a microbial community is overestimated. In addition to profiling both known (depicted in blue in the example below) and unknown (red) species represented in its MG database, mOTUs estimates the fraction of microbes in an environmental sample which are not represented in its genome collection, i.e the unassigned fraction (black).

Furthermore, mOTUs focuses on high precision, i.e. on detecting as many correct taxa (true positives) as possible while minimizing the detection of wrong ones (false positives). Together, these steps ensure a high quality of taxonomic profiles generated with mOTUs, which has been corroborated in independent benchmarks such as LEMMI and CAMI.
🗺️ Genomic Context: Access to genomes behind each mOTU
You can browse and query all 3.9 million genomes used for the construction of the mOTUs MG database
at mOTUs-db or using the motus genomes tool command (see tutorial).
The website allows you to search for mOTUs and genomes by taxonomic annotation, filter genomes by quality score, or find all MAGs reconstructed from a specific environmental sample or study.
The motus genomes command allows you to query genomes based on mOTUs and genome identifiers, taxonomic and
functional annotations. The output is directly compatible with motus download, which will download provided genome
sequences, enabling integration into custom pipelines.
🧩 User Genomes: Connect your genomes with existing mOTUs
By running the motus classify tool command, you can determine which mOTU does your genome of interest belong to,
place it in the context of processed mOTUs genomes and taxonomic profiles. Read our tutorial for more
information on how to do this.
⛓️ Long Reads: mOTUs profiles short as well as long reads
Long-read sequencing is becoming more and more popular for sequencing DNA from microbial communities or isolated strains.
There are few tools available for taxonomic profiling of long-read sequencing data, most of which rely on classifying and counting reads.
The long-read profiling pipeline in mOTUs performs reasonably well when compared with other pipelines specifically developed for long-read
sequencing data (see Portik et al., 2022).
You only need to run motus prep_long -i input_file -o output_file before running the regular profiling workflow.
⚡ Simple: Easy to install, lightweight and fast
You can easily install mOTUs using conda or pip. Once installed, you can obtain a taxonomic profile with a single shell command (see Quickstart). To increase accessibility, mOTUs3 also available as a QIIME2 plugin .
mOTUs is faster and requires less memory than most taxonomic profiling tools. The figure below shows data from the independent benchmark study

🛠️ Functional Context: Bridging Taxonomy and Function with mOTUs
A mOTU represents a species-level cluster defined by marker genes derived from one or more genomes. Using the motus genomes command, you can unlock the functional annotation associated with these genomes. This allows you to seamlessly map taxonomic abundance to metabolic potential, enabling a deeper understanding of the functional landscape within your microbial community.
 mOTUs is part of SIB's portfolio of open tools and databases.
mOTUs is part of SIB's portfolio of open tools and databases.
 mOTUs is part of the ELIXIR-CH Service Delivery Plan.
mOTUs is part of the ELIXIR-CH Service Delivery Plan.