Navigating the website#
mOTUs Table#
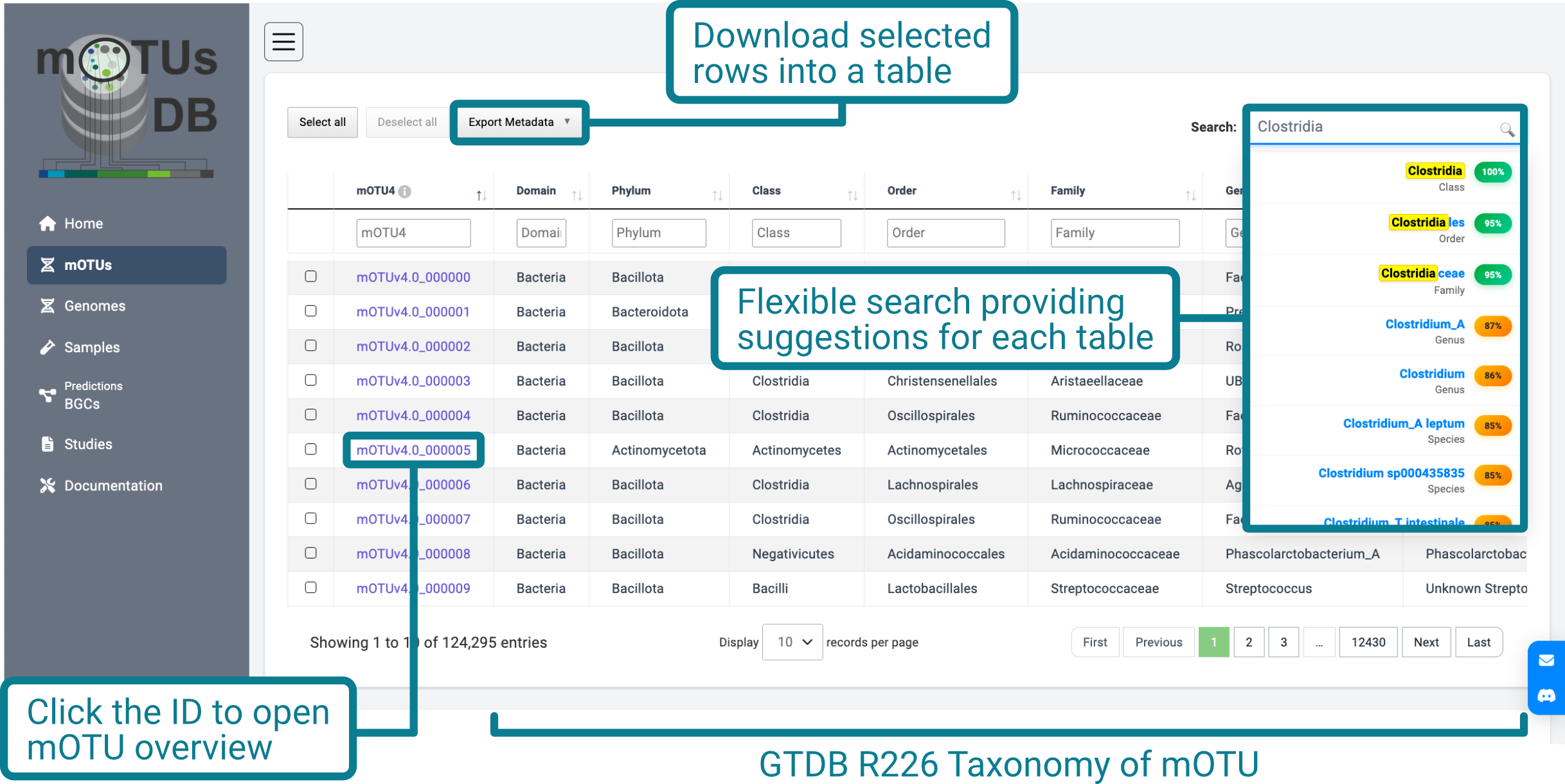
The following table provides a description of each column in the mOTU collection:
Field Name |
Type |
Description |
|---|---|---|
mOTU4 |
|
Unique identifier for the mOTU (compatible with versions 4.0 and 4.1). |
Domain |
|
Taxonomic domain assigned to the mOTU, derived from GTDB R226. |
Phylum |
|
Taxonomic phylum assigned to the mOTU, derived from GTDB R226. |
Class |
|
Taxonomic class assigned to the mOTU, derived from GTDB R226. |
Order |
|
Taxonomic order assigned to the mOTU, derived from GTDB R226. |
Family |
|
Taxonomic family assigned to the mOTU, derived from GTDB R226. |
Genus |
|
Taxonomic genus assigned to the mOTU, derived from GTDB R226. |
Species |
|
Taxonomic species assigned to the mOTU, derived from GTDB R226. |
MicrobeAtlas |
|
External link to the MicrobeAtlas database; returns a dash ( |
Representative |
|
Identifier for the genome designated as the high-quality cluster representative for this mOTU. |
Genome Table#
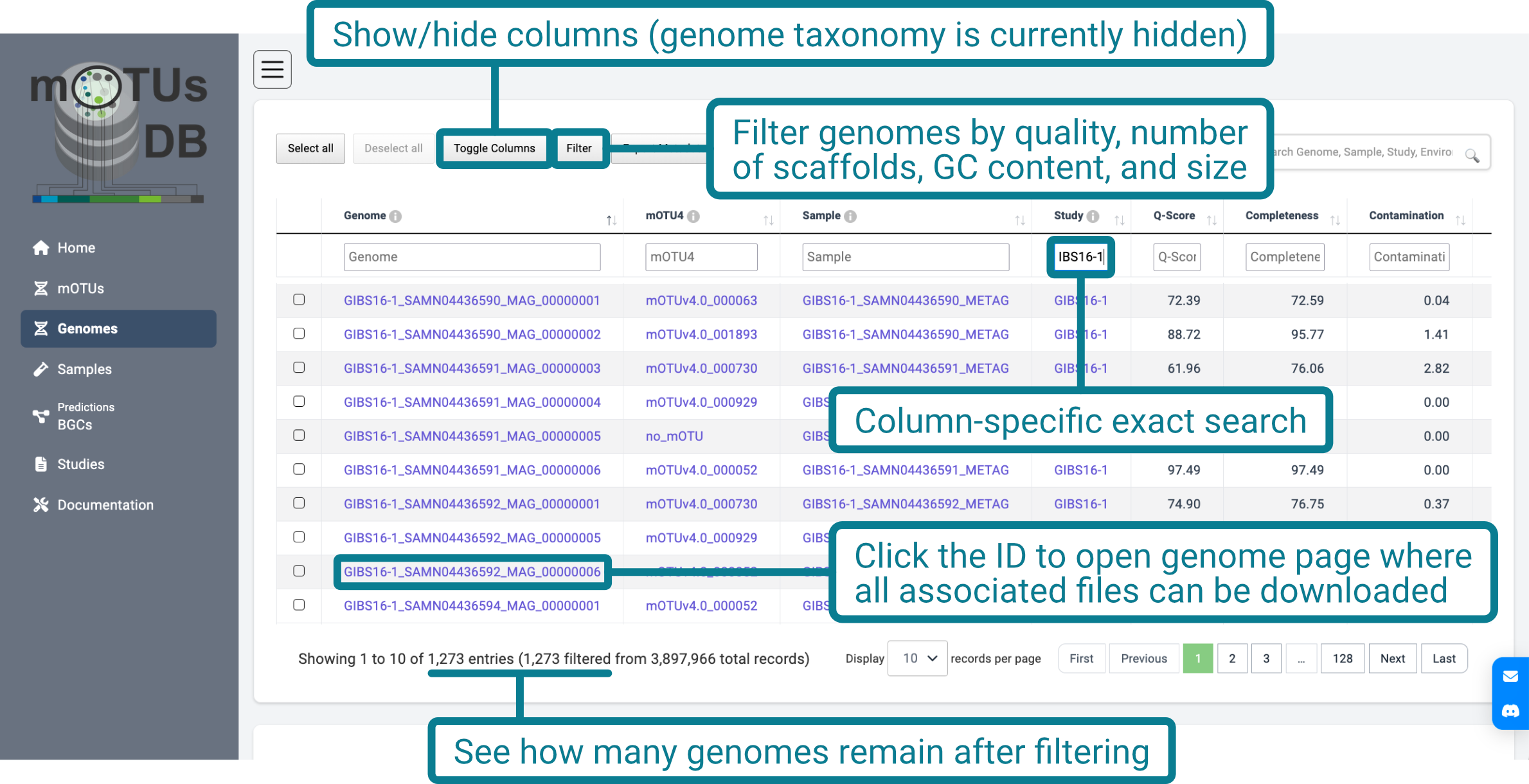
The following table defines the metadata fields and quality metrics associated with the individual genomes (MAGs, SAGs, and isolates) in the collection.
Field Name |
Data Type |
Description |
|---|---|---|
Genome |
|
Unique identifier or name of the individual genome. |
Domain |
|
Taxonomic domain derived from GTDB R226. |
Phylum |
|
Taxonomic phylum derived from GTDB R226. |
Class |
|
Taxonomic class derived from GTDB R226. |
Order |
|
Taxonomic order derived from GTDB R226. |
Family |
|
Taxonomic family derived from GTDB R226. |
Genus |
|
Taxonomic genus derived from GTDB R226. |
Species |
|
Taxonomic species derived from GTDB R226. |
mOTU4 |
|
The mOTU identifier (versions 4.0 and 4.1) to which this genome belongs. |
Sample |
|
The origin of the genomic data: the metagenomic sample for MAGs, or the source sample for Isolates/SAGs. |
Study |
|
The primary study or dataset from which the genome was recovered. |
Q-Score |
|
Integrated quality metric calculated as: \(Completeness - (5 \times Contamination)\). |
Completeness |
|
Estimated genome completeness percentage (0–100), determined by CheckM or Anvi’o. |
Contamination |
|
Estimated genomic contamination percentage (0–100), determined by CheckM or Anvi’o. |
N50 |
|
Assembly quality metric; the length of the shortest scaffold such that 50% of the total assembly length is contained in scaffolds of this size or larger. |
#Scaffolds |
|
The total number of scaffolds in the genome assembly. |
MAG |
|
Boolean flag; |
Is Rep |
|
Boolean flag; |
Representative |
|
The name of the genome designated as the cluster representative for the associated mOTU. |